|
12/27/2023 0 Comments Define okazaki fragment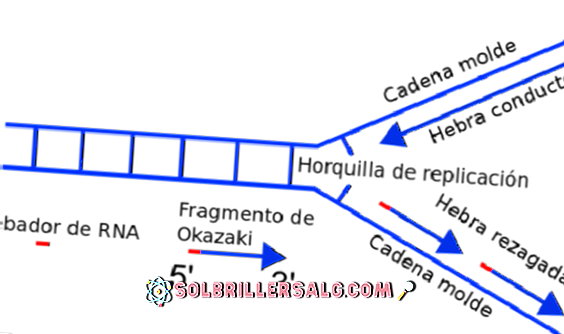
Small DNA substrates modeling the essence of the much larger lagging strand. The results reproduce experimental data, providing insights into events related to Okazaki fragment initiation and the overall functioning of DNA replisomes.ĭNA replication Okazaki fragment initiation T4 bacteriophage poymerase recycling stochastic modeling. We have modeled repeated cycles of Okazaki fragment initiation using a collision with a completed Okazaki fragment or primer-primase complexes as the recycling mechanism. The collision with primer-primase complexes triggering the early termination of Okazaki fragment synthesis has distinct advantages over those previously proposed because this signal requires no transmission to the lagging-strand polymerase through protein or DNA interactions, the mechanism for rapid dissociation of the holoenzyme is always collision, and no unique characteristics need to be assigned to either identical polymerase in the replisome. We show for the T4 bacteriophage DNA replication system that primer-primase complexes have a residence time similar to the timescale of Okazaki fragment synthesis and the ability to block a holoenzyme synthesizing DNA and stimulate the dissociation of the holoenzyme to trigger polymerase recycling. We examined the role of RNA primer-primase complexes left on the lagging ssDNA from primer synthesis in initiating early lagging-strand polymerase recycling. The mechanism and signal that initiate this behavior-that is, the signaling mechanism-have not been definitively identified. The lagging-strand polymerase sometimes recycles to begin the synthesis of a new Okazaki fragment before finishing the previous fragment, creating a gap between the Okazaki fragments. e.The opposite strand polarity of duplex DNA necessitates that the leading strand is replicated continuously whereas the lagging strand is replicated in discrete segments known as Okazaki fragments. It ensures proper processitvity of the polymerase, so it doesn't stop prematurely. It contains the active site for synthesis of RNA c. It recognizes promoters where transcription should begin. 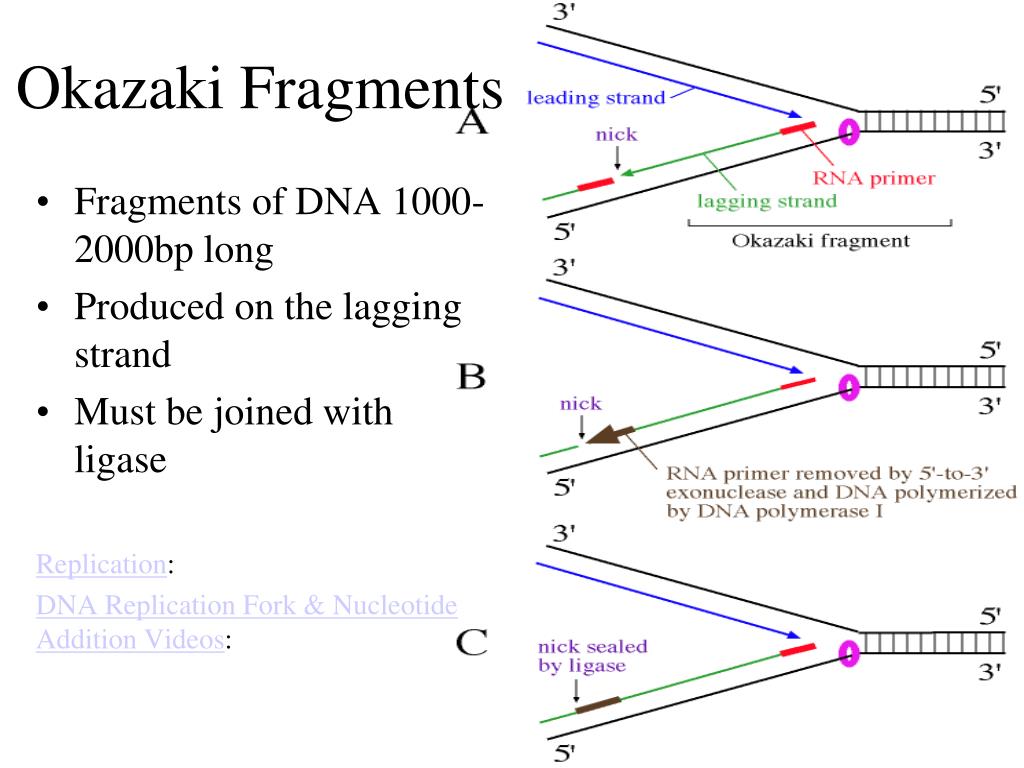
What is the function of the sigma (o) subunit of RNA polymerase in E. After RNA polymerase discovers an inverted repeat 15. Never it is an intrinsic part of the core enzyme e. After transcription begins and about 10 nucleotides have been added to the RNA chain. Prior to the incorporation of any nucleotides into an RNA strand. At what point does the sigma (?) subunit of RNA polymerase released from the core a. Which below is not a DNA strand used to direct RNA synthesis? (2 p) a. The core enzyme transcribes from an RNA template, the holoenzyme from a DNA template. The holoenzyme transerbes from an RNA template, the core enzyme from a DNA template. The core enzyme includes the sigma (o) subunit, the holoenzyme does not c. The holoenzyme includes the sigma (o) subunit, the core enzyme does not. How do the core enzyme and the holoenzyme of RNA polymerase differ in E. the remnants of the original strands that are dispersed in the new double stranded DNA molecules d. short DNA pieces that explain how DNA is synthesized on the leading strand. 
short DNA pieces that explain how DNA is synthesized on the lagging strand b.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
